Code
library(ggplot2)
library(data.table)
library(patchwork)
base_dir <- here::here()
source(file.path(base_dir, "sim-analyze.R"))
mcols <- method_colors()
mlabs <- method_labels()
dlabs <- dgp_short_labels()Registration Benchmark v2
study_b_keys <- c("method", "dgp", "noise_sd", "severity")
best <- aggregate_metric(
dt = dt,
metric = "warp_mise",
by = c(study_b_keys, "lambda"),
stat = "median",
value_name = "mise"
)
baseline <- best[lambda == 0, .(method, dgp, noise_sd, severity, base = mise)]
optimal <- find_optimal_by_metric(
dt = dt,
metric = "warp_mise",
by = study_b_keys,
option_col = "lambda",
stat = "median",
value_name = "mise"
)
warp_ratio_all <- ratio_to_baseline(
dt = dt,
metric = "warp_mise",
by = c(study_b_keys, "lambda"),
baseline_col = "lambda",
baseline_level = 0,
stat = "median",
value_name = "mise",
baseline_name = "base_mise",
ratio_name = "ratio",
keep_baseline = TRUE
)
merged <- merge(
optimal,
warp_ratio_all[, .(method, dgp, noise_sd, severity, lambda, base_mise, ratio)],
by = c("method", "dgp", "noise_sd", "severity", "lambda")
)
ae_best <- aggregate_metric(
dt = dt,
metric = "alignment_error",
by = c(study_b_keys, "lambda"),
stat = "median",
value_name = "ae"
)
ae_baseline <- ae_best[lambda == 0, .(method, dgp, noise_sd, severity, base = ae)]
ae_optimal <- find_optimal_by_metric(
dt = dt,
metric = "alignment_error",
by = study_b_keys,
option_col = "lambda",
stat = "median",
value_name = "ae"
)
ae_ratio_all <- ratio_to_baseline(
dt = dt,
metric = "alignment_error",
by = c(study_b_keys, "lambda"),
baseline_col = "lambda",
baseline_level = 0,
stat = "median",
value_name = "ae",
baseline_name = "base_ae",
ratio_name = "ratio",
keep_baseline = TRUE
)
ae_merged <- merge(
ae_optimal,
ae_ratio_all[, .(method, dgp, noise_sd, severity, lambda, base_ae, ratio)],
by = c("method", "dgp", "noise_sd", "severity", "lambda")
)
lambda_comp <- compare_optima(
opt_a = optimal,
opt_b = ae_optimal,
key_cols = study_b_keys,
value_cols = c("lambda", "lambda"),
value_names = c("mise_lambda", "ae_lambda")
)Study B tests whether regularization (lambda > 0) can rescue registration methods in regimes where they struggle. Design: 4 DGPs \(\times\) 8 lambdas \(\times\) 3 methods (SRVF, CC default, CC crit1) \(\times\) 3 noise \(\times\) 2 severity \(\times\) 33 reps = 19,008 runs.
Lambda grid calibrated by pilot (job 5131464): c(0, 1e-4, 1e-3, 0.01, 0.05, 0.1, 1, 10).
Conditional-on-success MISE comparisons can be biased if failure rates vary by lambda. The table below checks this.
fail_by_lambda <- dt[,
.(n_total = .N,
n_fail = sum(failure),
rate_pct = sprintf("%.1f", 100 * mean(failure))),
by = .(method, lambda)
]
fail_wide <- dcast(fail_by_lambda,
method ~ lambda,
value.var = "rate_pct")
knitr::kable(fail_wide,
caption = "Failure rate (%) by method and lambda, pooling across DGPs, noise, severity, and reps.")| method | 0 | 1e-04 | 0.001 | 0.01 | 0.05 | 0.1 | 1 | 10 |
|---|---|---|---|---|---|---|---|---|
| srvf | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 |
| cc_default | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 |
| cc_crit1 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 |
The core question: does the MISE-vs-lambda curve show a U-shape (optimal lambda > 0)?
agg <- dt[failure == FALSE,
.(mise = median(warp_mise, na.rm = TRUE),
q25 = quantile(warp_mise, 0.25, na.rm = TRUE),
q75 = quantile(warp_mise, 0.75, na.rm = TRUE)),
by = .(method, dgp, noise_sd, severity, lambda)
]
# Baseline (lambda = 0) for reference lines
baseline <- agg[lambda == 0, .(method, dgp, noise_sd, severity, base = mise)]
ggplot(agg[severity == "1"],
aes(x = lambda + 1e-5, y = mise, color = method)) +
geom_line(linewidth = 0.6) +
geom_point(size = 1.5) +
geom_errorbar(aes(ymin = q25, ymax = q75), width = 0.15, alpha = 0.4) +
geom_hline(data = baseline[severity == "1"],
aes(yintercept = base, color = method),
linetype = "dashed", alpha = 0.5) +
facet_grid(dgp ~ noise_sd,
scales = "free_y",
labeller = labeller(
noise_sd = function(x) paste0("noise = ", x),
dgp = dlabs)) +
scale_x_log10(
breaks = c(1e-5, 1e-4, 1e-3, 0.01, 0.05, 0.1, 1, 10),
labels = c("0", "1e-4", "1e-3", "0.01", "0.05", "0.1", "1", "10")) +
scale_color_manual(values = mcols, labels = mlabs) +
scale_y_log10() +
labs(x = expression(lambda ~ "(log scale)"),
y = "Warp MISE (median, log scale)",
color = "Method") +
theme_benchmark() +
theme(legend.position = "bottom")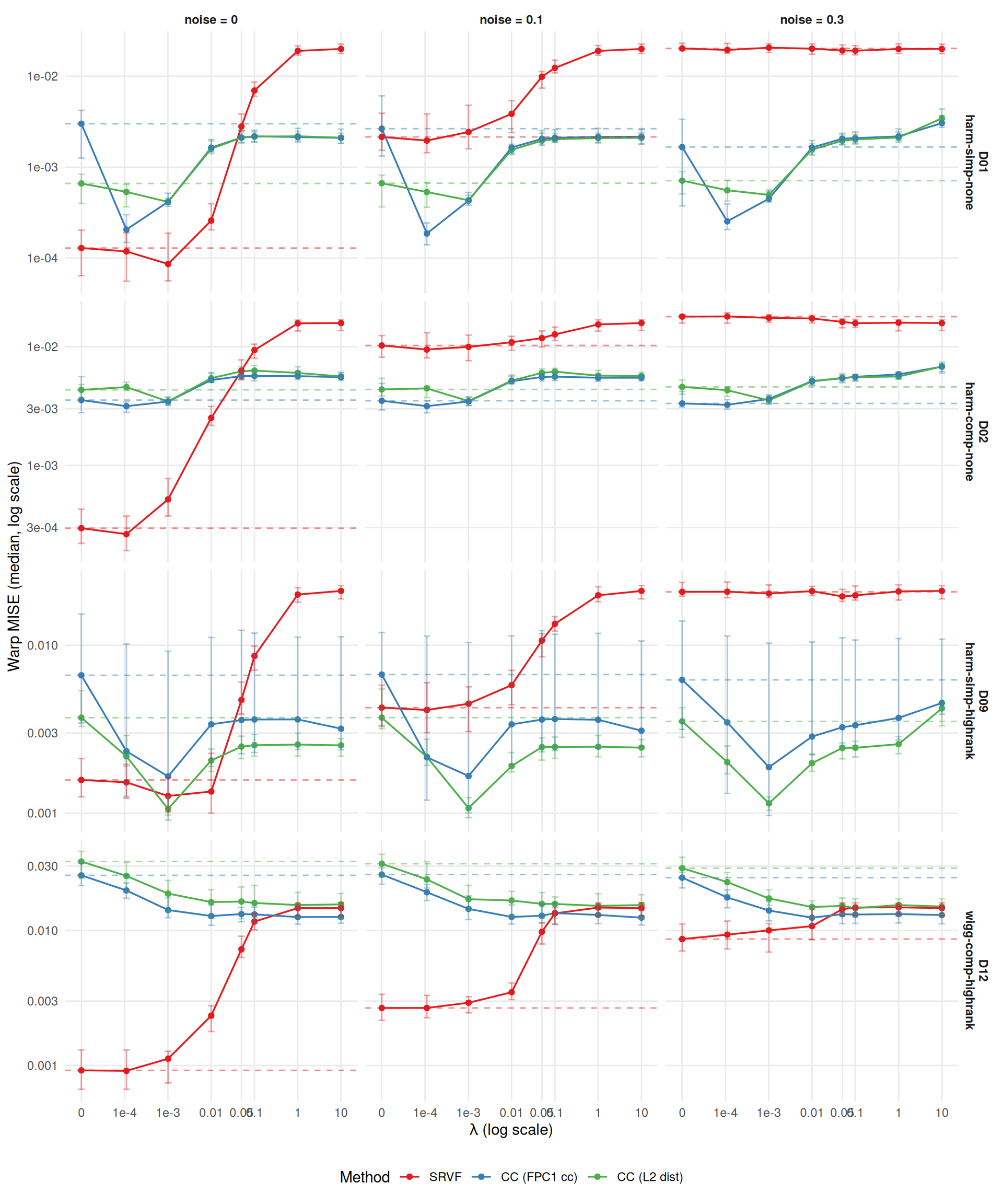
ggplot(agg[severity == "0.5"],
aes(x = lambda + 1e-5, y = mise, color = method)) +
geom_line(linewidth = 0.6) +
geom_point(size = 1.5) +
geom_errorbar(aes(ymin = q25, ymax = q75), width = 0.15, alpha = 0.4) +
geom_hline(data = baseline[severity == "0.5"],
aes(yintercept = base, color = method),
linetype = "dashed", alpha = 0.5) +
facet_grid(dgp ~ noise_sd,
scales = "free_y",
labeller = labeller(
noise_sd = function(x) paste0("noise = ", x),
dgp = dlabs)) +
scale_x_log10(
breaks = c(1e-5, 1e-4, 1e-3, 0.01, 0.05, 0.1, 1, 10),
labels = c("0", "1e-4", "1e-3", "0.01", "0.05", "0.1", "1", "10")) +
scale_color_manual(values = mcols, labels = mlabs) +
scale_y_log10() +
labs(x = expression(lambda ~ "(log scale)"),
y = "Warp MISE (median, log scale)",
color = "Method") +
theme_benchmark() +
theme(legend.position = "bottom")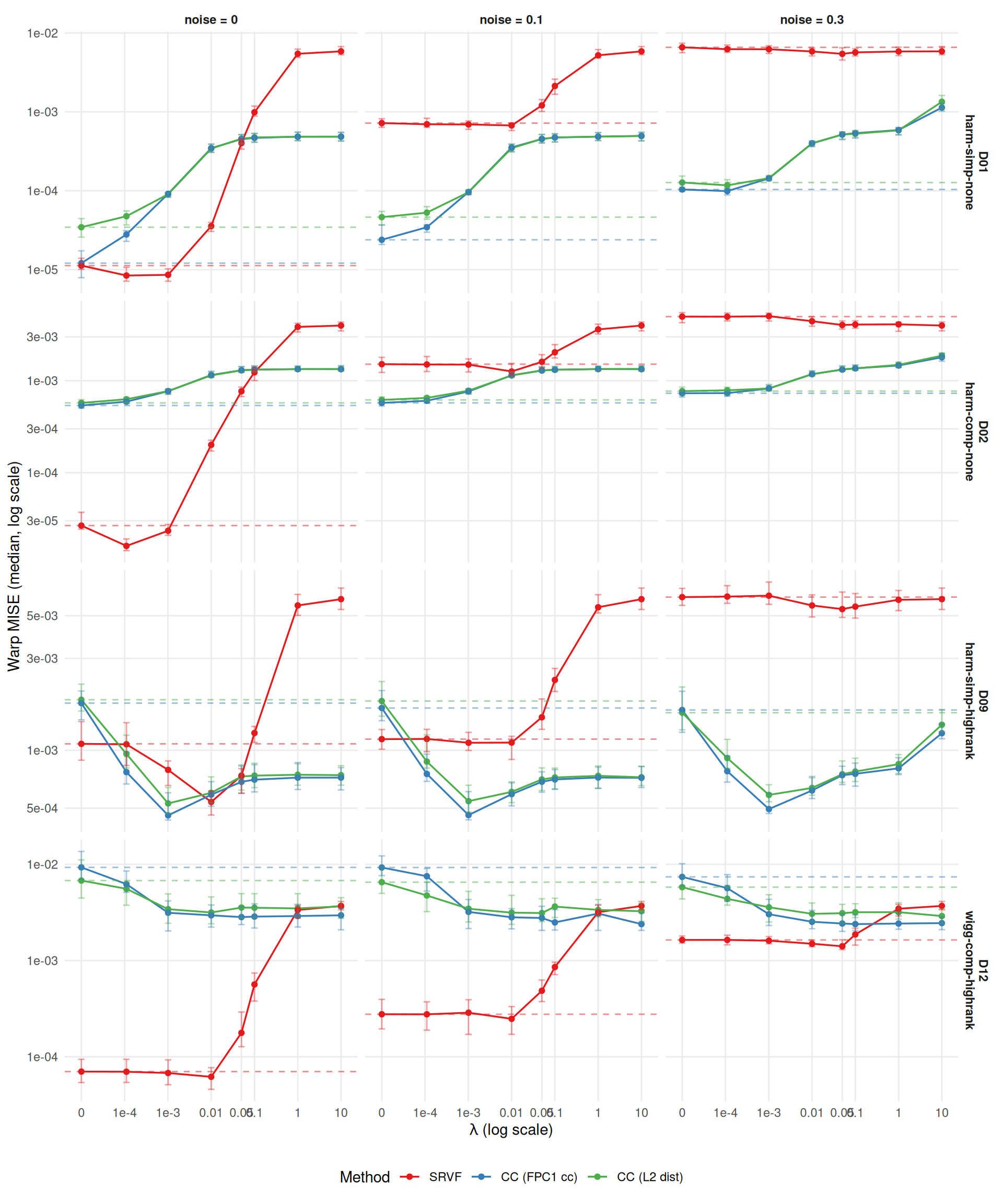
| method | dgp | severity | 0 | 0.1 | 0.3 |
|---|---|---|---|---|---|
| srvf | D01 | 0.5 | 0.000 | 1e-02 | 5e-02 |
| srvf | D01 | 1 | 0.001 | 0e+00 | 1e-01 |
| srvf | D02 | 0.5 | 0.000 | 1e-02 | 1e+01 |
| srvf | D02 | 1 | 0.000 | 0e+00 | 1e-01 |
| srvf | D09 | 0.5 | 0.010 | 1e-03 | 5e-02 |
| srvf | D09 | 1 | 0.001 | 0e+00 | 5e-02 |
| srvf | D12 | 0.5 | 0.010 | 1e-02 | 5e-02 |
| srvf | D12 | 1 | 0.000 | 0e+00 | 0e+00 |
| cc_default | D01 | 0.5 | 0.000 | 0e+00 | 0e+00 |
| cc_default | D01 | 1 | 0.000 | 0e+00 | 0e+00 |
| cc_default | D02 | 0.5 | 0.000 | 0e+00 | 0e+00 |
| cc_default | D02 | 1 | 0.000 | 0e+00 | 0e+00 |
| cc_default | D09 | 0.5 | 0.001 | 1e-03 | 1e-03 |
| cc_default | D09 | 1 | 0.001 | 1e-03 | 1e-03 |
| cc_default | D12 | 0.5 | 0.050 | 1e+01 | 1e-01 |
| cc_default | D12 | 1 | 1.000 | 1e+01 | 1e-02 |
| cc_crit1 | D01 | 0.5 | 0.000 | 0e+00 | 0e+00 |
| cc_crit1 | D01 | 1 | 0.001 | 1e-03 | 1e-03 |
| cc_crit1 | D02 | 0.5 | 0.000 | 0e+00 | 0e+00 |
| cc_crit1 | D02 | 1 | 0.001 | 1e-03 | 1e-03 |
| cc_crit1 | D09 | 0.5 | 0.001 | 1e-03 | 1e-03 |
| cc_crit1 | D09 | 1 | 0.001 | 1e-03 | 1e-03 |
| cc_crit1 | D12 | 0.5 | 0.010 | 5e-02 | 1e+01 |
| cc_crit1 | D12 | 1 | 1.000 | 1e+00 | 1e-01 |
for (m in levels(dt$method)) {
sub <- optimal[method == m]
# Most common optimal lambda per noise level (mode, not median — grid is discrete)
by_noise <- sub[, .(
mode_lambda = {
tab <- table(lambda)
as.numeric(names(tab)[which.max(tab)])
},
n_zero = sum(lambda == 0),
n_total = .N
), by = noise_sd]
cat(sprintf("- **%s**: most common optimal λ by noise: %s. ",
mlabs[m],
paste(sprintf("noise=%s → %g (%d/%d at λ=0)",
by_noise$noise_sd, by_noise$mode_lambda,
by_noise$n_zero, by_noise$n_total), collapse = "; ")))
# Full frequency table across all conditions
freq <- sub[, .N, by = lambda][order(lambda)]
freq_txt <- paste(sprintf("%g: %d", freq$lambda, freq$N), collapse = ", ")
cat(sprintf("Distribution: {%s}. ", freq_txt))
# Flag boundary hits
n_boundary <- sum(sub$lambda == max(sub$lambda))
if (n_boundary > 0) {
cat(sprintf("**%d/%d cells hit boundary (λ=%g).**",
n_boundary, nrow(sub), max(sub$lambda)))
}
cat("\n")
}How much does the best lambda improve over lambda=0?
Note: This is an oracle analysis — the optimal lambda is selected ex post from the same data used to evaluate it. Ratios \(\leq 1\) by construction (the baseline lambda=0 is always in the candidate set). The practical benefit of penalization depends on whether a good lambda can be selected a priori or via cross-validation.
min_ratio <- min(merged$ratio, na.rm = TRUE)
ggplot(merged,
aes(x = interaction(noise_sd, severity, sep = " / sev="),
y = interaction(dgp, method, sep = " : "),
fill = ratio)) +
geom_tile(color = "white") +
geom_text(aes(label = sprintf("%.2f\n(λ=%g)", ratio, lambda)),
size = 2.5) +
scale_fill_gradient(
low = "#2ca02c", high = "white",
limits = c(min_ratio, 1),
oob = scales::squish
) +
labs(x = "noise / severity",
y = "DGP : Method",
fill = "MISE ratio\n(vs λ=0)") +
theme_benchmark() +
theme(axis.text.x = element_text(angle = 45, hjust = 1))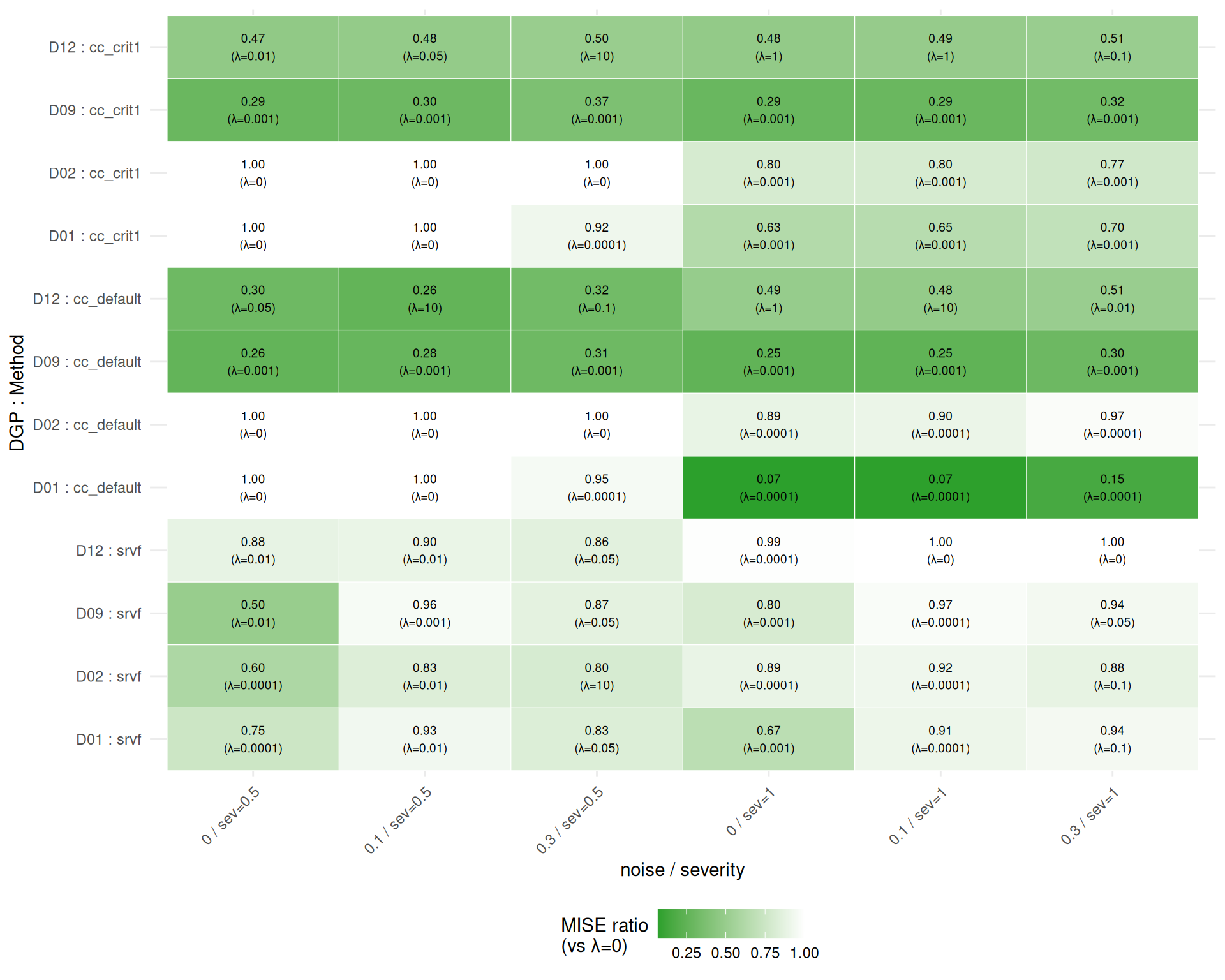
Findings:
Average improvement ratio across the 24 method-specific cells (mean ± SE; lower is better):
# Biggest winners
big_win <- merged[ratio < 0.8][order(ratio)]
if (nrow(big_win) > 0) {
cat("Cases where penalization improves MISE by >20%:\n\n")
for (i in seq_len(min(nrow(big_win), 10))) {
r <- big_win[i]
cat(sprintf("- %s on %s (noise=%s, sev=%s): %.0f%% improvement at λ=%g\n",
mlabs[r$method], r$dgp, r$noise_sd, r$severity,
100 * (1 - r$ratio), r$lambda))
}
}Cases where penalization improves MISE by >20%:
Does cc_crit1 respond differently to lambda than cc_default?
# Check whether severity=0.5 patterns differ qualitatively
fda_both <- dt[failure == FALSE & method %in% c("cc_default", "cc_crit1")]
fda_agg_both <- fda_both[,
.(mise = median(warp_mise, na.rm = TRUE)),
by = .(method, dgp, noise_sd, severity, lambda)
]
# Correlation of MISE profiles across severity
cor_default <- cor(
fda_agg_both[method == "cc_default" & severity == "0.5", mise],
fda_agg_both[method == "cc_default" & severity == "1", mise],
use = "complete.obs"
)
cor_crit1 <- cor(
fda_agg_both[method == "cc_crit1" & severity == "0.5", mise],
fda_agg_both[method == "cc_crit1" & severity == "1", mise],
use = "complete.obs"
)
cat(sprintf(
"Results shown for severity=1.0. MISE profiles are highly correlated across severities (r=%.2f for cc_default, r=%.2f for cc_crit1); severity=0.5 patterns are qualitatively similar.\n\n",
cor_default, cor_crit1))Results shown for severity=1.0. MISE profiles are highly correlated across severities (r=0.95 for cc_default, r=0.99 for cc_crit1); severity=0.5 patterns are qualitatively similar.
fda_dt <- dt[failure == FALSE & method %in% c("cc_default", "cc_crit1")]
fda_agg <- aggregate_median_iqr(
dt = fda_dt[severity == "1"],
metric = "warp_mise",
by = c("method", "dgp", "noise_sd", "lambda"),
value_name = "mise"
)
ggplot(fda_agg,
aes(x = lambda + 1e-5, y = mise, color = method)) +
geom_line(linewidth = 0.8) +
geom_point(size = 2) +
geom_errorbar(aes(ymin = q25, ymax = q75), width = 0.15, alpha = 0.35) +
facet_grid(dgp ~ noise_sd,
scales = "free_y",
labeller = labeller(
noise_sd = function(x) paste0("noise = ", x),
dgp = dlabs)) +
scale_x_log10(
breaks = c(1e-5, 1e-4, 1e-3, 0.01, 0.05, 0.1, 1, 10),
labels = c("0", "1e-4", "1e-3", "0.01", "0.05", "0.1", "1", "10")) +
scale_color_manual(values = mcols, labels = mlabs) +
scale_y_log10() +
labs(x = expression(lambda ~ "(log scale)"),
y = "Warp MISE (median, log scale)",
color = "Method") +
theme_benchmark() +
theme(legend.position = "bottom")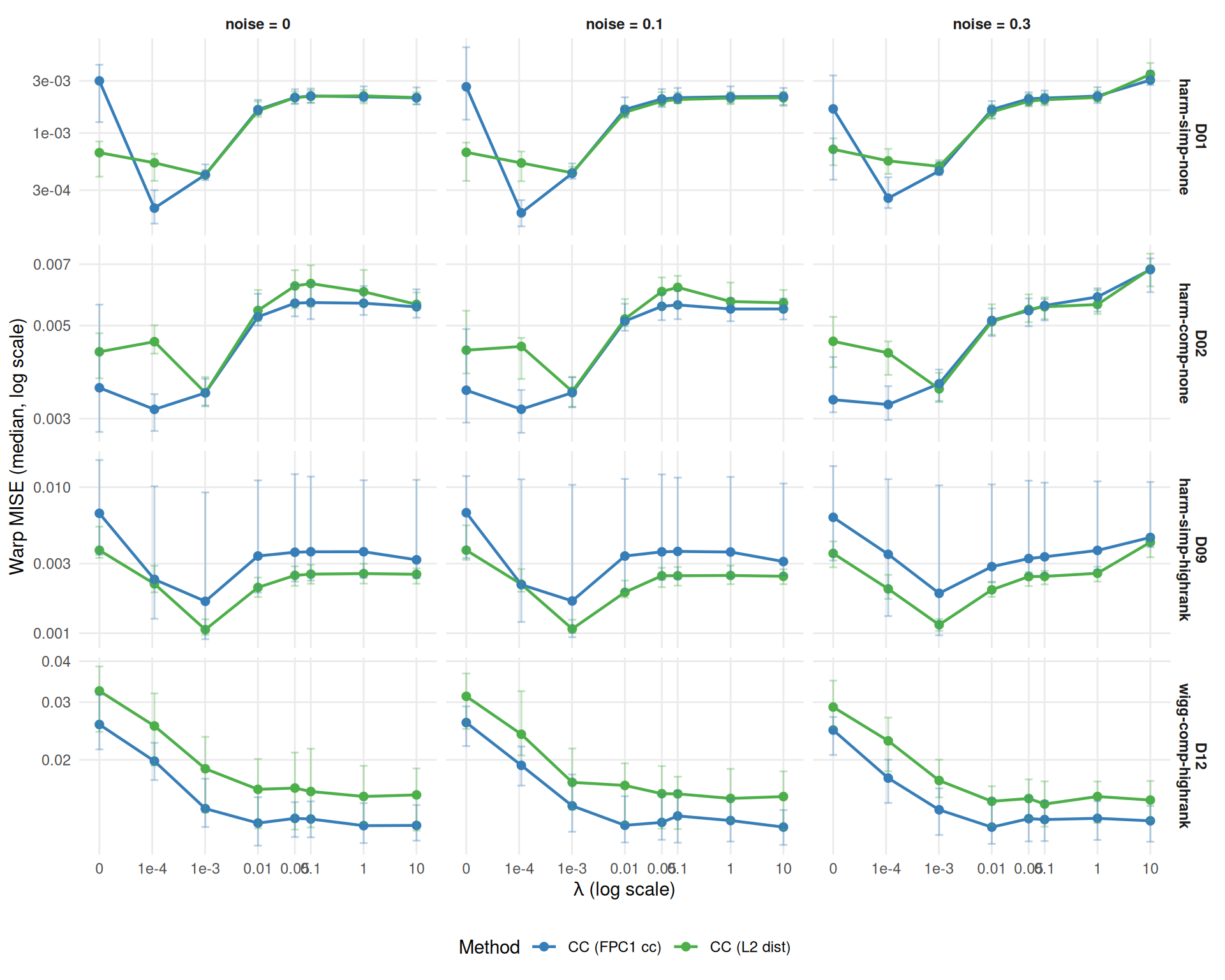
# Compare optimal lambdas for the two CC methods
fda_opt <- optimal[method %in% c("cc_default", "cc_crit1")]
fda_comp <- dcast(
fda_opt[, .(method, dgp, noise_sd, severity, lambda, mise)],
dgp + noise_sd + severity ~ method,
value.var = list("lambda", "mise")
)
cat("**CC criterion comparison:**\n\n")CC criterion comparison:
# Paired comparison: how often is crit1's optimal lambda higher/lower/equal?
fda_comp[, crit1_higher := lambda_cc_crit1 > lambda_cc_default]
fda_comp[, crit1_lower := lambda_cc_crit1 < lambda_cc_default]
fda_comp[, crit1_equal := lambda_cc_crit1 == lambda_cc_default]
n_higher <- sum(fda_comp$crit1_higher, na.rm = TRUE)
n_lower <- sum(fda_comp$crit1_lower, na.rm = TRUE)
n_equal <- sum(fda_comp$crit1_equal, na.rm = TRUE)
n_comp <- nrow(fda_comp)
cat(sprintf(
"- Paired comparison of optimal λ: cc_crit1 higher in %d/%d, lower in %d/%d, equal in %d/%d cells.\n",
n_higher, n_comp, n_lower, n_comp, n_equal, n_comp))SRVF’s lambda response is expected to differ qualitatively from CC methods.
srvf_both <- dt[failure == FALSE & method == "srvf"]
srvf_agg_both <- srvf_both[,
.(mise = median(warp_mise, na.rm = TRUE)),
by = .(dgp, noise_sd, severity, lambda)
]
cor_srvf <- cor(
srvf_agg_both[severity == "0.5", mise],
srvf_agg_both[severity == "1", mise],
use = "complete.obs"
)
cat(sprintf(
"Results shown for severity=1.0. SRVF MISE profiles across severities are highly correlated (r=%.2f); severity=0.5 patterns are qualitatively similar.\n\n",
cor_srvf))Results shown for severity=1.0. SRVF MISE profiles across severities are highly correlated (r=0.94); severity=0.5 patterns are qualitatively similar.
srvf_agg <- dt[failure == FALSE & method == "srvf" & severity == "1",
.(mise = median(warp_mise, na.rm = TRUE),
q25 = quantile(warp_mise, 0.25, na.rm = TRUE),
q75 = quantile(warp_mise, 0.75, na.rm = TRUE)),
by = .(dgp, noise_sd, lambda)
]
ggplot(srvf_agg,
aes(x = lambda + 1e-5, y = mise, color = noise_sd)) +
geom_line(linewidth = 0.8) +
geom_point(size = 2) +
geom_ribbon(aes(ymin = q25, ymax = q75, fill = noise_sd),
alpha = 0.15, color = NA) +
facet_wrap(~dgp, scales = "free_y", labeller = as_labeller(dlabs)) +
scale_x_log10(
breaks = c(1e-5, 1e-4, 1e-3, 0.01, 0.05, 0.1, 1, 10),
labels = c("0", "1e-4", "1e-3", "0.01", "0.05", "0.1", "1", "10")) +
scale_color_brewer(palette = "Set1") +
scale_fill_brewer(palette = "Set1") +
scale_y_log10() +
labs(x = expression(lambda ~ "(log scale)"),
y = "Warp MISE (median ± IQR, log scale)",
color = "Noise", fill = "Noise") +
theme_benchmark() +
theme(legend.position = "bottom")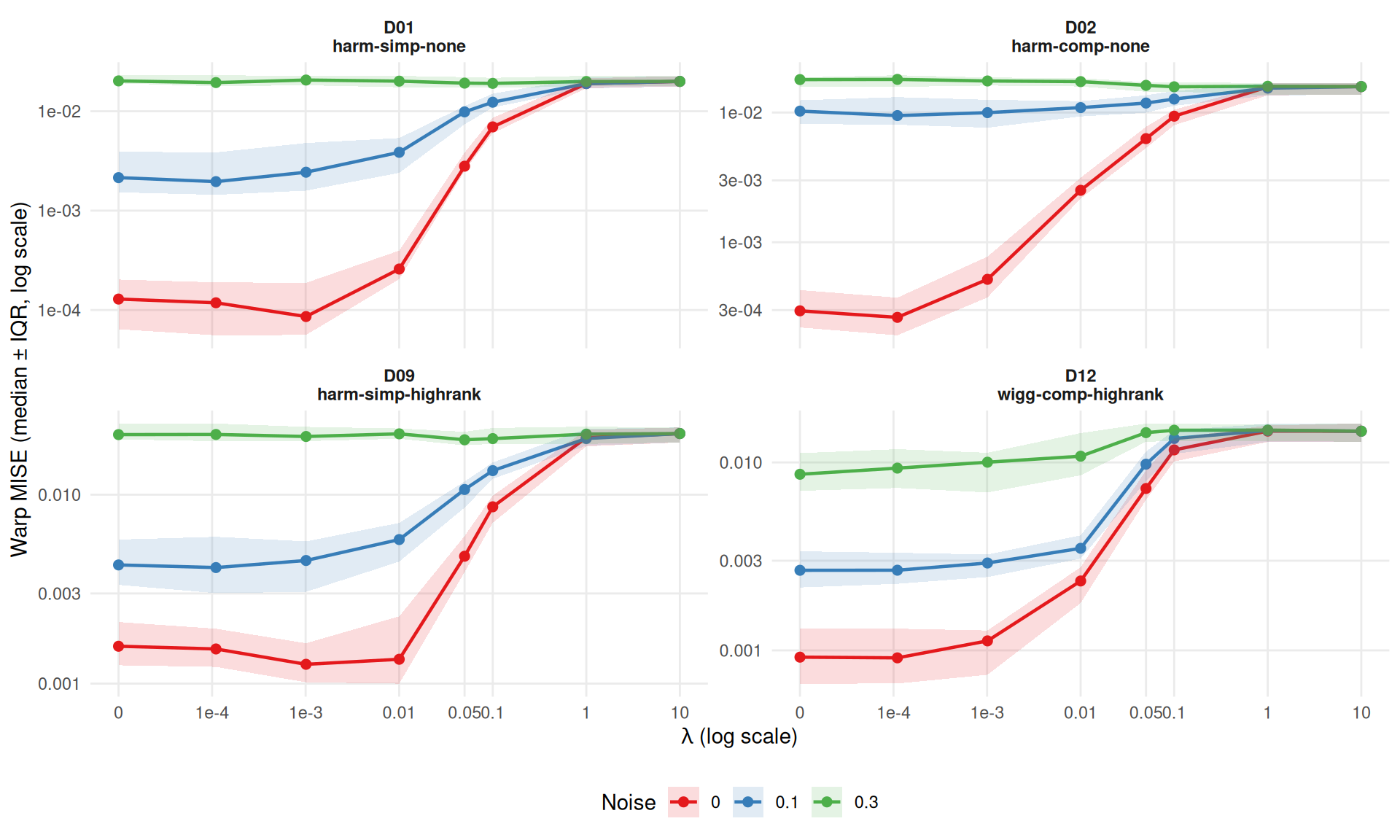
SRVF lambda findings:
# Does lambda ever help SRVF substantially?
srvf_merged <- merged[method == "srvf"]
helps <- srvf_merged[ratio < 0.9]
if (nrow(helps) > 0) {
cat(sprintf("Lambda helps SRVF (>10%% improvement) in %d/%d conditions:\n",
nrow(helps), nrow(srvf_merged)))
for (i in seq_len(nrow(helps))) {
r <- helps[i]
cat(sprintf(" - %s noise=%s sev=%s: %.0f%% at λ=%g\n",
r$dgp, r$noise_sd, r$severity, 100*(1-r$ratio), r$lambda))
}
} else {
cat("Lambda does not substantially help SRVF (>10%) in any condition.\n")
}Lambda helps SRVF (>10% improvement) in 14/24 conditions: - D01 noise=0 sev=0.5: 25% at λ=0.0001 - D01 noise=0 sev=1: 33% at λ=0.001 - D01 noise=0.3 sev=0.5: 17% at λ=0.05 - D02 noise=0 sev=0.5: 40% at λ=0.0001 - D02 noise=0 sev=1: 11% at λ=0.0001 - D02 noise=0.1 sev=0.5: 17% at λ=0.01 - D02 noise=0.3 sev=0.5: 20% at λ=10 - D02 noise=0.3 sev=1: 12% at λ=0.1 - D09 noise=0 sev=0.5: 50% at λ=0.01 - D09 noise=0 sev=1: 20% at λ=0.001 - D09 noise=0.3 sev=0.5: 13% at λ=0.05 - D12 noise=0 sev=0.5: 12% at λ=0.01 - D12 noise=0.1 sev=0.5: 10% at λ=0.01 - D12 noise=0.3 sev=0.5: 14% at λ=0.05
Average SRVF improvement ratio across cells: 0.86 ± 0.03 (n=24).
Does lambda affect template estimation quality?
tmpl_both <- dt[failure == FALSE,
.(tmise = median(template_mise, na.rm = TRUE)),
by = .(method, dgp, noise_sd, severity, lambda)
]
cor_tmpl <- cor(
tmpl_both[severity == "0.5", tmise],
tmpl_both[severity == "1", tmise],
use = "complete.obs"
)
cat(sprintf(
"Results shown for severity=1.0. Template MISE profiles across severities are highly correlated (r=%.2f); severity=0.5 patterns are qualitatively similar.\n\n",
cor_tmpl))Results shown for severity=1.0. Template MISE profiles across severities are highly correlated (r=0.87); severity=0.5 patterns are qualitatively similar.
tmpl_agg <- aggregate_median_iqr(
dt = dt[failure == FALSE & severity == "1"],
metric = "template_mise",
by = c("method", "dgp", "noise_sd", "lambda"),
value_name = "tmise"
)
ggplot(tmpl_agg,
aes(x = lambda + 1e-5, y = tmise, color = method)) +
geom_line(linewidth = 0.6) +
geom_point(size = 1.5) +
geom_errorbar(aes(ymin = q25, ymax = q75), width = 0.15, alpha = 0.35) +
facet_grid(dgp ~ noise_sd,
scales = "free_y",
labeller = labeller(
noise_sd = function(x) paste0("noise = ", x),
dgp = dlabs)) +
scale_x_log10(
breaks = c(1e-5, 1e-4, 1e-3, 0.01, 0.05, 0.1, 1, 10),
labels = c("0", "1e-4", "1e-3", "0.01", "0.05", "0.1", "1", "10")) +
scale_color_manual(values = mcols, labels = mlabs) +
scale_y_log10() +
labs(x = expression(lambda ~ "(log scale)"),
y = "Template MISE (median, log scale)",
color = "Method") +
theme_benchmark() +
theme(legend.position = "bottom")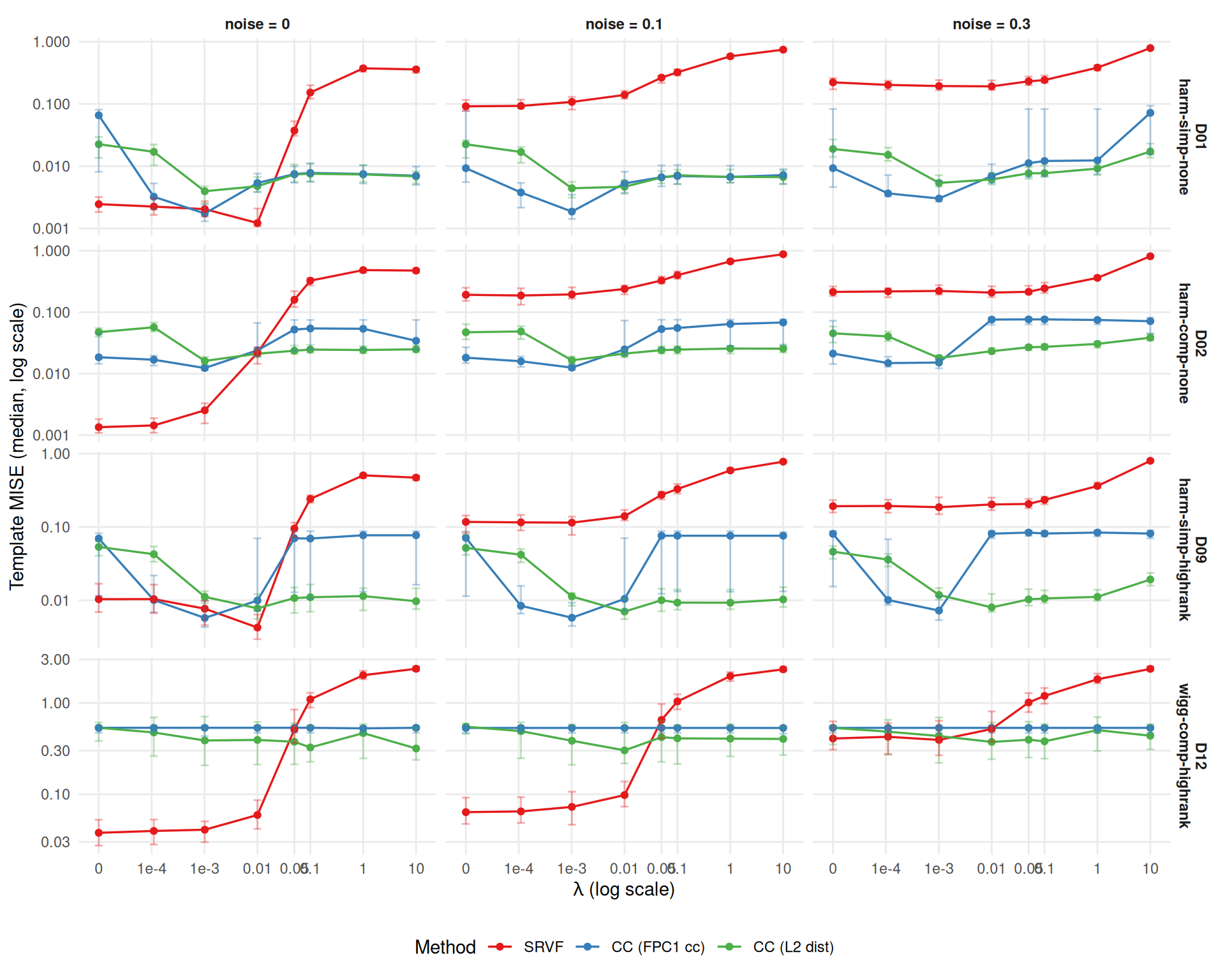
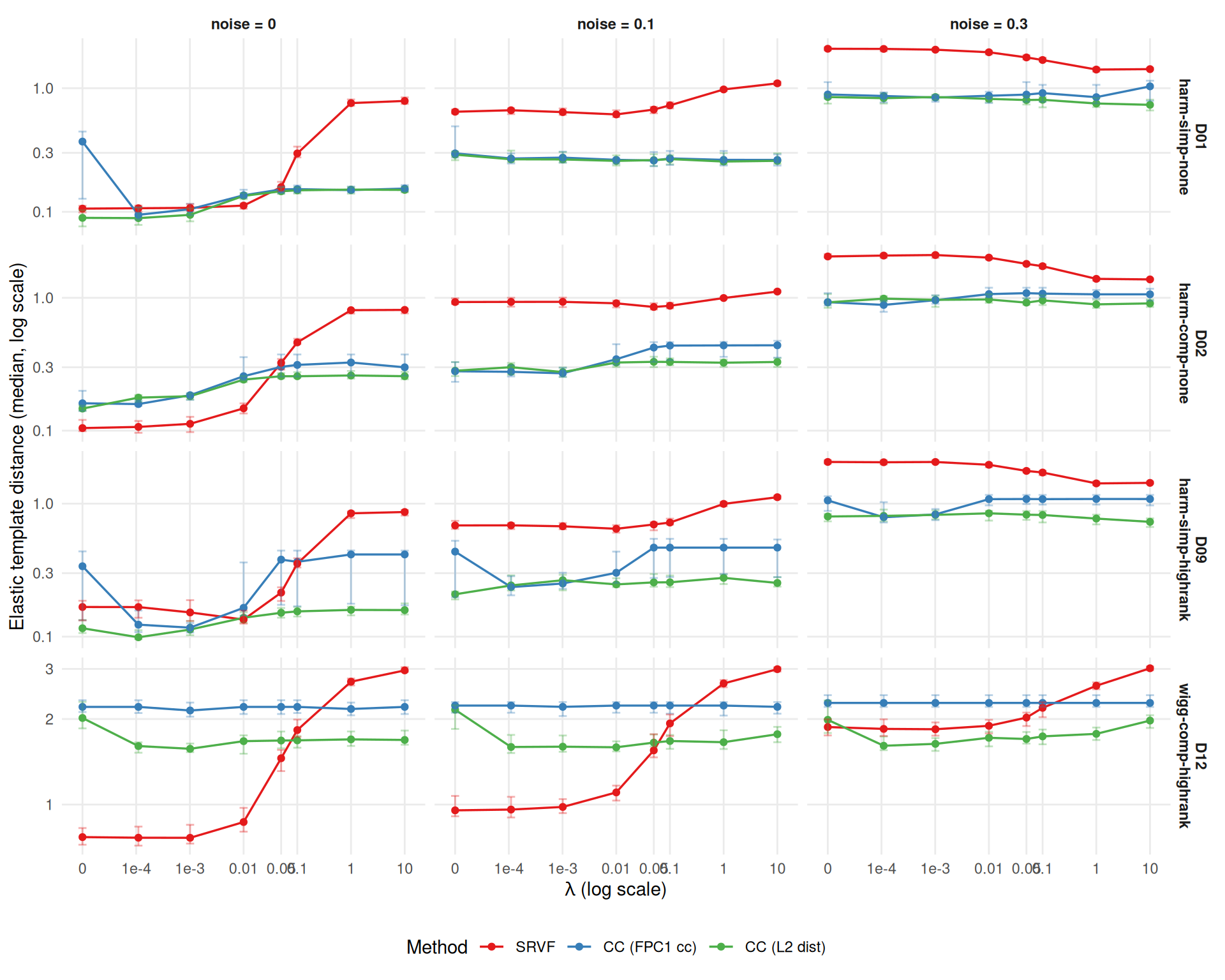
has_elastic_b <- "template_elastic_dist" %in% names(dt) &&
any(!is.na(dt$template_elastic_dist))
if (has_elastic_b) {
edist_agg <- aggregate_median_iqr(
dt = dt[failure == FALSE & severity == "1" &
!is.na(template_elastic_dist)],
metric = "template_elastic_dist",
by = c("method", "dgp", "noise_sd", "lambda"),
value_name = "edist"
)
ggplot(edist_agg,
aes(x = lambda + 1e-5, y = edist, color = method)) +
geom_line(linewidth = 0.6) +
geom_point(size = 1.5) +
geom_errorbar(aes(ymin = q25, ymax = q75), width = 0.15, alpha = 0.35) +
facet_grid(dgp ~ noise_sd,
scales = "free_y",
labeller = labeller(
noise_sd = function(x) paste0("noise = ", x),
dgp = dlabs)) +
scale_x_log10(
breaks = c(1e-5, 1e-4, 1e-3, 0.01, 0.05, 0.1, 1, 10),
labels = c("0", "1e-4", "1e-3", "0.01", "0.05", "0.1", "1", "10")) +
scale_color_manual(values = mcols, labels = mlabs) +
scale_y_log10() +
labs(x = expression(lambda ~ "(log scale)"),
y = "Elastic template distance (median, log scale)",
color = "Method") +
theme_benchmark() +
theme(legend.position = "bottom")
} else {
cat("*Elastic template distance not available in Study B results.*")
}
if (has_elastic_b) {
# Optimize each template metric separately to avoid mixed-target bias
tmpl_base <- dt[failure == FALSE & lambda == 0 & severity == "1",
.(tmise_base = median(template_mise, na.rm = TRUE),
edist_base = median(template_elastic_dist, na.rm = TRUE)),
by = .(method, dgp, noise_sd)]
tmpl_by_lambda <- dt[failure == FALSE & severity == "1" &
!is.na(template_elastic_dist),
.(tmise = median(template_mise, na.rm = TRUE),
edist = median(template_elastic_dist, na.rm = TRUE)),
by = .(method, dgp, noise_sd, lambda)]
# Best lambda for elastic distance
best_edist <- tmpl_by_lambda[,
.SD[which.min(edist)],
by = .(method, dgp, noise_sd)]
edist_comp <- merge(best_edist, tmpl_base, by = c("method", "dgp", "noise_sd"))
edist_comp[, edist_ratio := edist / edist_base]
# Best lambda for template MISE (optimized independently)
best_tmise <- tmpl_by_lambda[,
.SD[which.min(tmise)],
by = .(method, dgp, noise_sd)]
tmise_comp <- merge(best_tmise, tmpl_base, by = c("method", "dgp", "noise_sd"))
tmise_comp[, tmise_ratio := tmise / tmise_base]
n_edist_helps <- sum(edist_comp$edist_ratio < 0.9)
n_tmise_helps <- sum(tmise_comp$tmise_ratio < 0.9)
cat(sprintf(
"**Template quality under penalization** (each metric optimized at its own best λ): elastic template distance improves >10%% in %d/%d cells; template MISE improves >10%% in %d/%d cells. ",
n_edist_helps, nrow(edist_comp), n_tmise_helps, nrow(tmise_comp)))
# Which method benefits most for template shape?
avg_edist <- edist_comp[, .(mean_ratio = mean(edist_ratio, na.rm = TRUE)),
by = method]
best_tmpl_m <- avg_edist[which.min(mean_ratio)]
edist_method_summary <- mean_se_table(edist_comp, "edist_ratio", "method")
best_tmpl_summary <- edist_method_summary[method == best_tmpl_m$method]
cat(sprintf(
"%s benefits most for template *shape* (mean elastic distance ratio %s).\n\n",
mlabs[best_tmpl_m$method],
fmt_mean_se(best_tmpl_summary$mean, best_tmpl_summary$se)
))
}Template quality under penalization (each metric optimized at its own best λ): elastic template distance improves >10% in 15/36 cells; template MISE improves >10% in 24/36 cells. CC (FPC1 cc) benefits most for template shape (mean elastic distance ratio 0.80 ± 0.08).
Alignment error measures how well registered curves match the true pre-warp curves: \(\text{mean}_i \int (\hat{x}_i^{\text{aligned}} - \tilde{x}_i)^2\,dt\). This captures the practical downstream impact of registration quality, complementing the warp MISE.
ae_agg <- dt[failure == FALSE,
.(ae = median(alignment_error, na.rm = TRUE),
q25 = quantile(alignment_error, 0.25, na.rm = TRUE),
q75 = quantile(alignment_error, 0.75, na.rm = TRUE)),
by = .(method, dgp, noise_sd, severity, lambda)
]
ae_baseline <- ae_agg[lambda == 0, .(method, dgp, noise_sd, severity, base = ae)]
ggplot(ae_agg[severity == "1"],
aes(x = lambda + 1e-5, y = ae, color = method)) +
geom_line(linewidth = 0.6) +
geom_point(size = 1.5) +
geom_errorbar(aes(ymin = q25, ymax = q75), width = 0.15, alpha = 0.4) +
geom_hline(data = ae_baseline[severity == "1"],
aes(yintercept = base, color = method),
linetype = "dashed", alpha = 0.5) +
facet_grid(dgp ~ noise_sd,
scales = "free_y",
labeller = labeller(
noise_sd = function(x) paste0("noise = ", x),
dgp = dlabs)) +
scale_x_log10(
breaks = c(1e-5, 1e-4, 1e-3, 0.01, 0.05, 0.1, 1, 10),
labels = c("0", "1e-4", "1e-3", "0.01", "0.05", "0.1", "1", "10")) +
scale_color_manual(values = mcols, labels = mlabs) +
scale_y_log10() +
labs(x = expression(lambda ~ "(log scale)"),
y = "Alignment error (median, log scale)",
color = "Method") +
theme_benchmark() +
theme(legend.position = "bottom")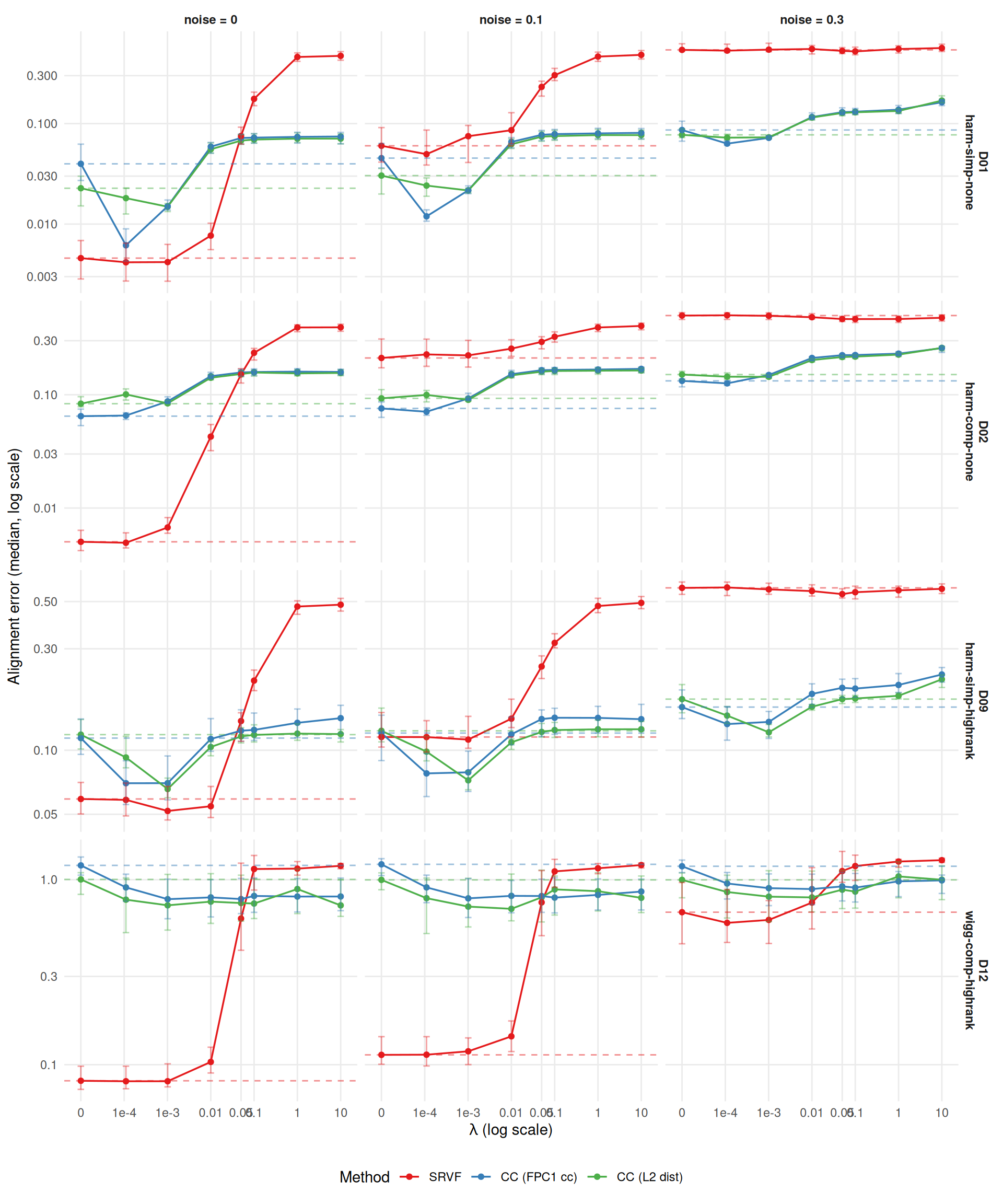
Alignment error: optimal lambda and improvement
# Correlation between alignment error and warp MISE improvement ratios
warp_merged_sub <- merged[, .(method, dgp, noise_sd, severity, warp_ratio = ratio)]
ae_merged_sub <- ae_merged[, .(method, dgp, noise_sd, severity, ae_ratio = ratio)]
both <- merge(warp_merged_sub, ae_merged_sub,
by = c("method", "dgp", "noise_sd", "severity"))
r_improvement <- cor(both$warp_ratio, both$ae_ratio, use = "complete.obs")
cat(sprintf("- Correlation between warp MISE and alignment error improvement ratios: r = %.2f.\n",
r_improvement))Average alignment-error improvement ratio across the 24 method-specific cells (mean ± SE):
# Biggest improvement
biggest_ae <- ae_merged[which.min(ratio)]
cat(sprintf("- Largest alignment error improvement: %s on %s (noise=%s, sev=%s): %.0f%% at λ=%g.\n\n",
mlabs[biggest_ae$method], biggest_ae$dgp, biggest_ae$noise_sd, biggest_ae$severity,
100 * (1 - biggest_ae$ratio), biggest_ae$lambda))if (r_improvement > 0.9) {
cat("The two metrics respond nearly identically to penalization — the same lambdas ",
"that improve warp recovery also improve alignment quality.\n")
} else if (r_improvement > 0.7) {
cat("The metrics are strongly correlated but not identical — in some cells, the ",
"lambda that minimizes warp MISE is not optimal for alignment error.\n")
} else {
cat("The metrics show moderate agreement — penalization affects warp recovery ",
"and alignment quality somewhat differently.\n")
}The metrics are strongly correlated but not identical — in some cells, the lambda that minimizes warp MISE is not optimal for alignment error.
# Summary table: best lambda and improvement per method x DGP
summary_dt <- copy(merged)
summary_dt[, improvement := sprintf("%.0f%%", 100 * (1 - mise / base_mise))]
summary_dt[, cell := sprintf("λ=%g (%s)", lambda, improvement)]
summary_wide <- dcast(
summary_dt[severity == "1", .(method, dgp, noise_sd, cell)],
method + dgp ~ noise_sd,
value.var = "cell"
)
cat("## Optimal lambda and improvement at severity=1.0\n\n")| method | dgp | 0 | 0.1 | 0.3 |
|---|---|---|---|---|
| srvf | D01 | λ=0.001 (33%) | λ=0.0001 (9%) | λ=0.1 (6%) |
| srvf | D02 | λ=0.0001 (11%) | λ=0.0001 (8%) | λ=0.1 (12%) |
| srvf | D09 | λ=0.001 (20%) | λ=0.0001 (3%) | λ=0.05 (6%) |
| srvf | D12 | λ=0.0001 (1%) | λ=0 (0%) | λ=0 (0%) |
| cc_default | D01 | λ=0.0001 (93%) | λ=0.0001 (93%) | λ=0.0001 (85%) |
| cc_default | D02 | λ=0.0001 (11%) | λ=0.0001 (10%) | λ=0.0001 (3%) |
| cc_default | D09 | λ=0.001 (75%) | λ=0.001 (75%) | λ=0.001 (70%) |
| cc_default | D12 | λ=1 (51%) | λ=10 (52%) | λ=0.01 (49%) |
| cc_crit1 | D01 | λ=0.001 (37%) | λ=0.001 (35%) | λ=0.001 (30%) |
| cc_crit1 | D02 | λ=0.001 (20%) | λ=0.001 (20%) | λ=0.001 (23%) |
| cc_crit1 | D09 | λ=0.001 (71%) | λ=0.001 (71%) | λ=0.001 (68%) |
| cc_crit1 | D12 | λ=1 (52%) | λ=1 (51%) | λ=0.1 (49%) |
Note: statistics below pool both severities unless stated otherwise.