if (has_elastic) {
# Rank correlation between the two metrics
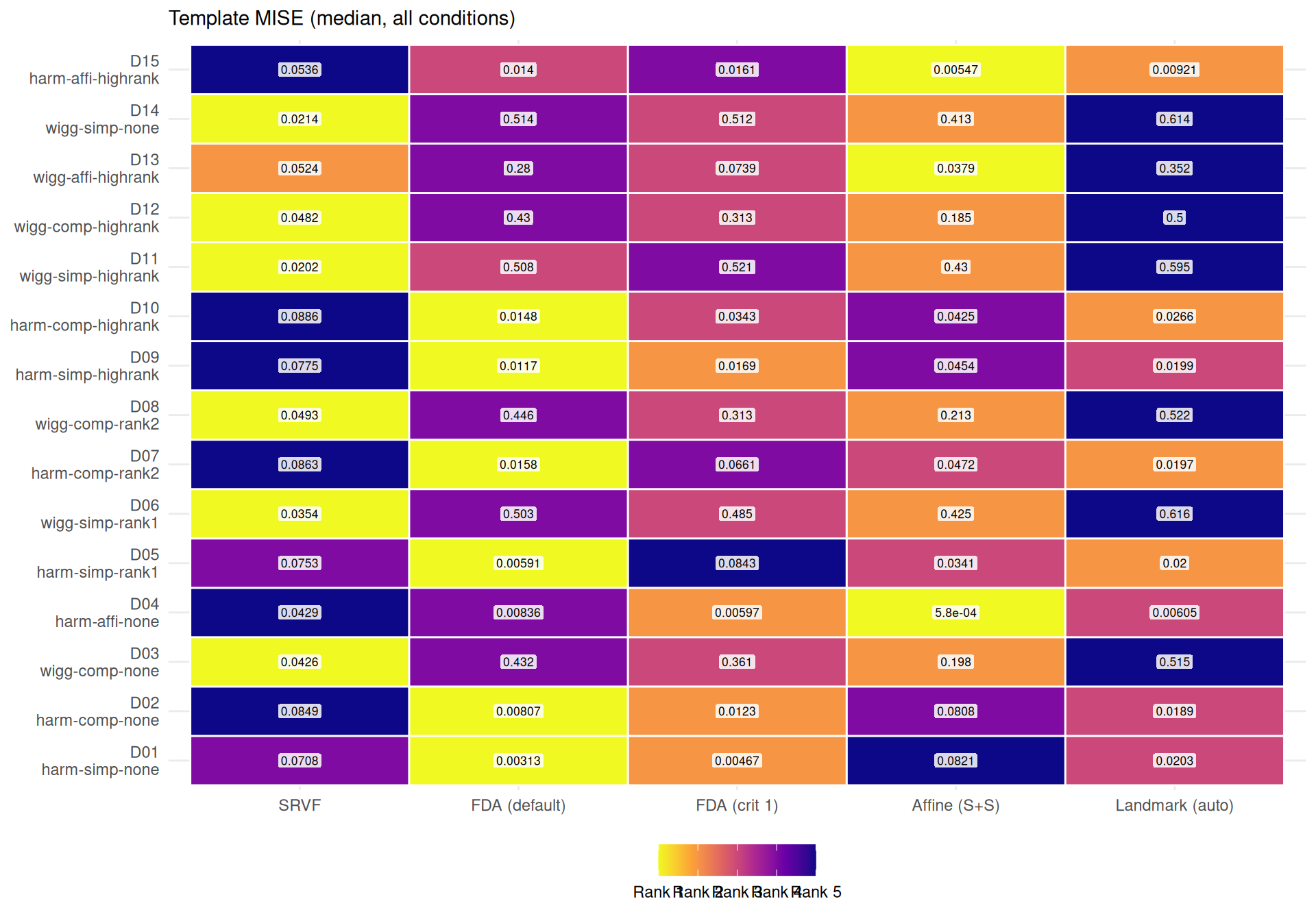
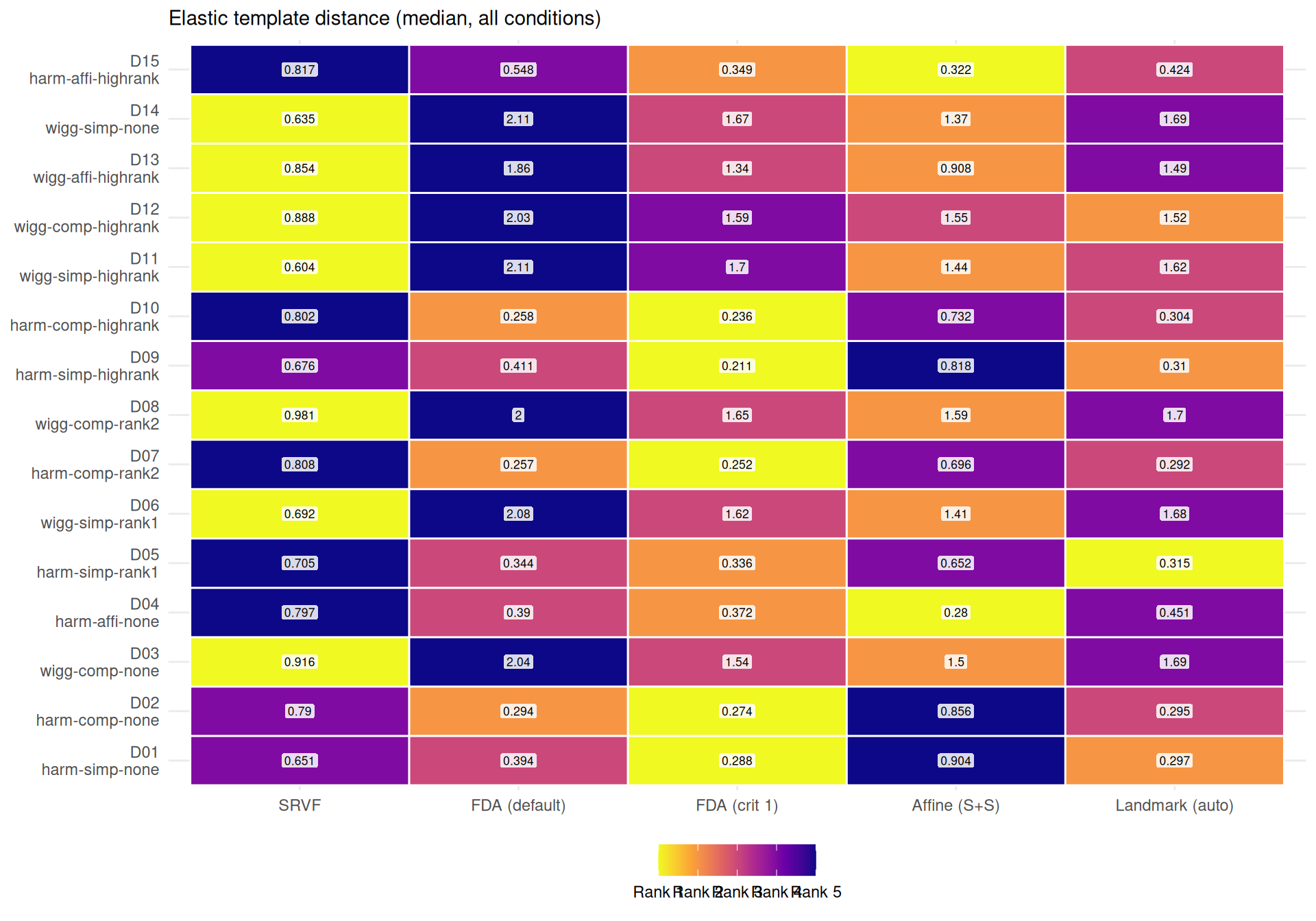
agg_both <- dt[failure == FALSE & !is.na(template_elastic_dist) & !is.na(template_mise),
.(edist = median(template_elastic_dist), tmise = median(template_mise)),
by = .(dgp, severity, noise_sd, method)]
agg_both[, rank_e := frank(edist), by = .(dgp, severity, noise_sd)]
agg_both[, rank_m := frank(tmise), by = .(dgp, severity, noise_sd)]
rho <- round(cor(agg_both$rank_e, agg_both$rank_m, method = "spearman"), 2)
# Best-method disagreement
best_e <- agg_both[, .SD[which.min(edist)], by = .(dgp, severity, noise_sd)]
best_m <- agg_both[, .SD[which.min(tmise)], by = .(dgp, severity, noise_sd)]
merged <- merge(
best_e[, .(dgp, severity, noise_sd, best_elastic = method)],
best_m[, .(dgp, severity, noise_sd, best_mise = method)]
)
n_disagree <- sum(merged$best_elastic != merged$best_mise)
n_total <- nrow(merged)
# Win rates by elastic distance
agg_both[, rank_e2 := frank(edist), by = .(dgp, severity, noise_sd)]
wr <- agg_both[, .(win_pct = round(100 * mean(rank_e2 == 1), 1)), by = method][order(-win_pct)]
# Noise sensitivity ratio (noise=0.3 / noise=0), conditioned on template
noise_sens <- dt[
failure == FALSE & !is.na(template_elastic_dist) & noise_sd %in% c("0", "0.3"),
.(edist = median(template_elastic_dist)),
by = .(method, template, noise_sd)
]
wide <- dcast(noise_sens, method + template ~ noise_sd, value.var = "edist")
setnames(wide, c("0", "0.3"), c("e0", "e03"))
wide[, ratio := round(e03 / e0, 2)]
# Summarize across templates (median of template-specific ratios)
wide_pooled <- wide[, .(ratio = round(median(ratio), 1)), by = method]
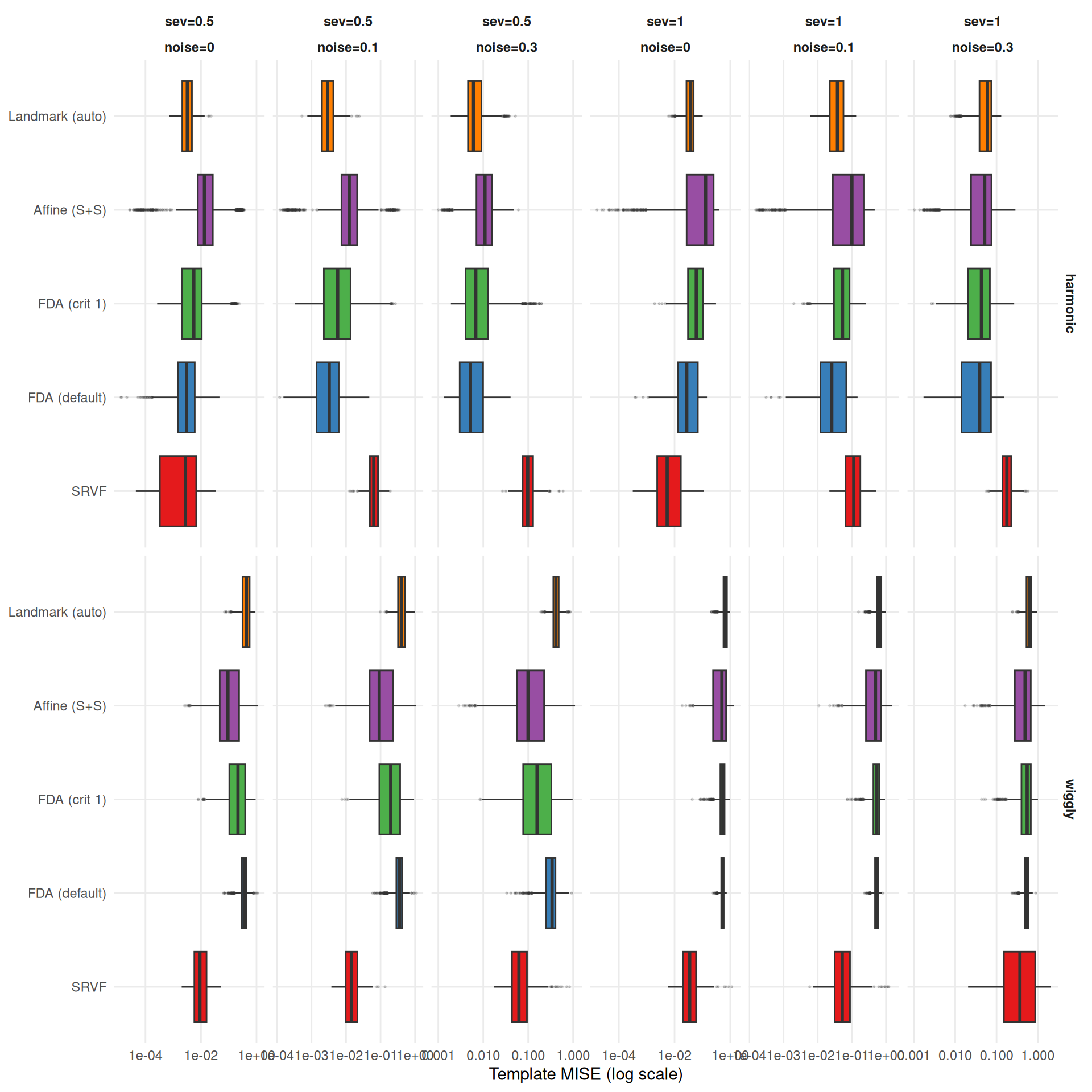
cat(sprintf(
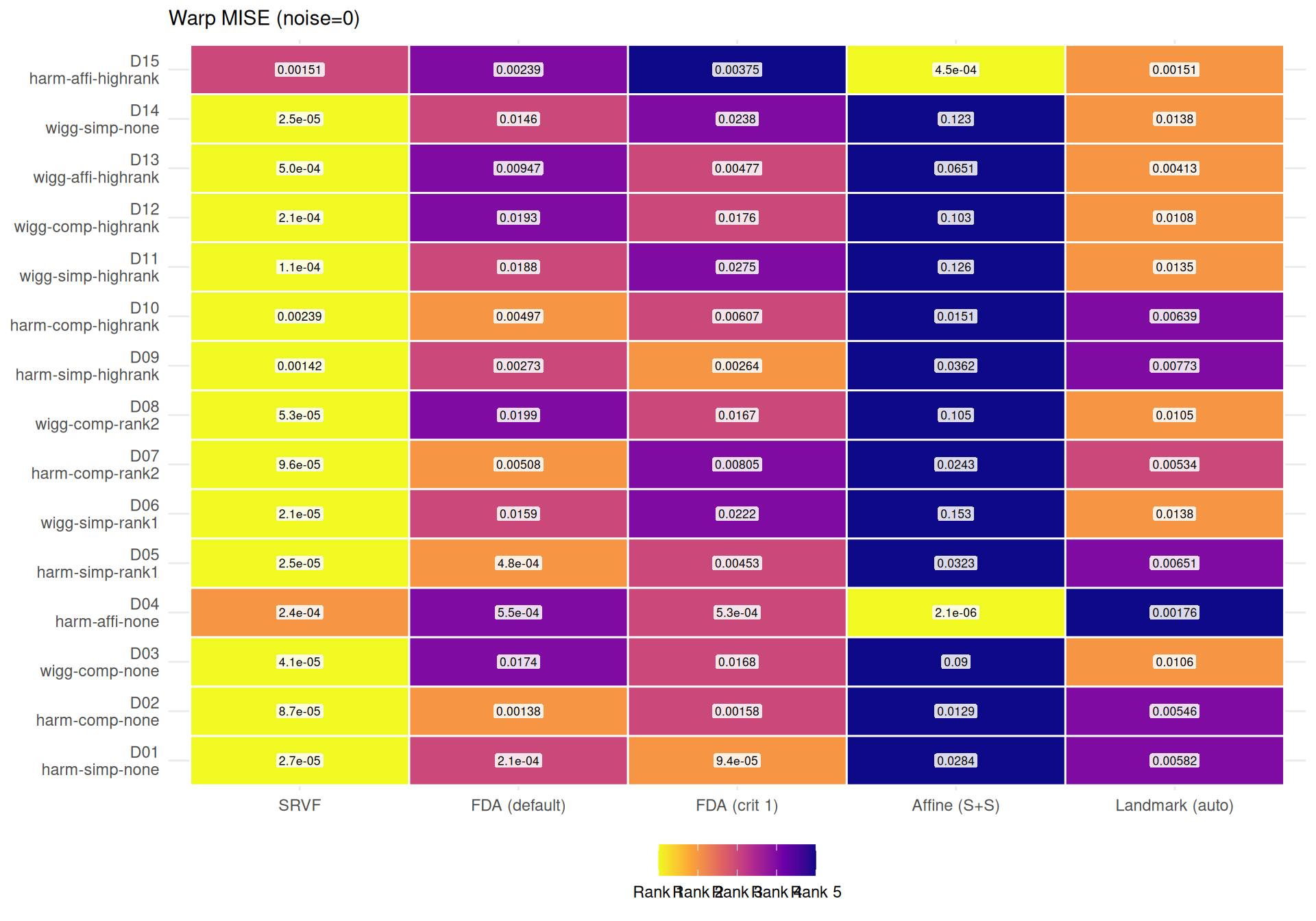
"The two metrics agree moderately (Spearman $\\rho$ = %s across cell-level method rankings) but disagree on the best method in **%d of %d** conditions (%.0f%%). Key differences:\n\n",
rho, n_disagree, n_total, 100 * n_disagree / n_total
))
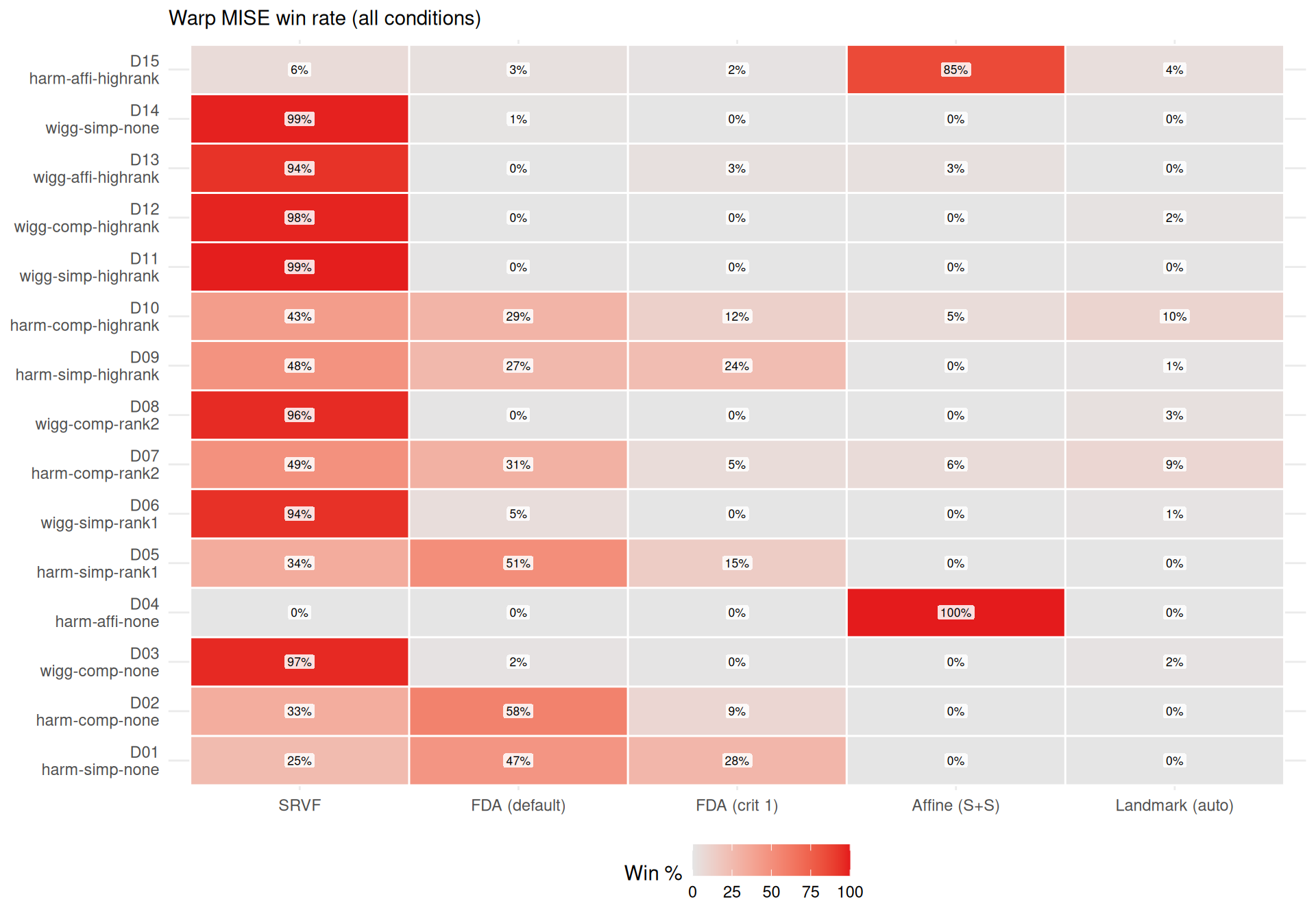
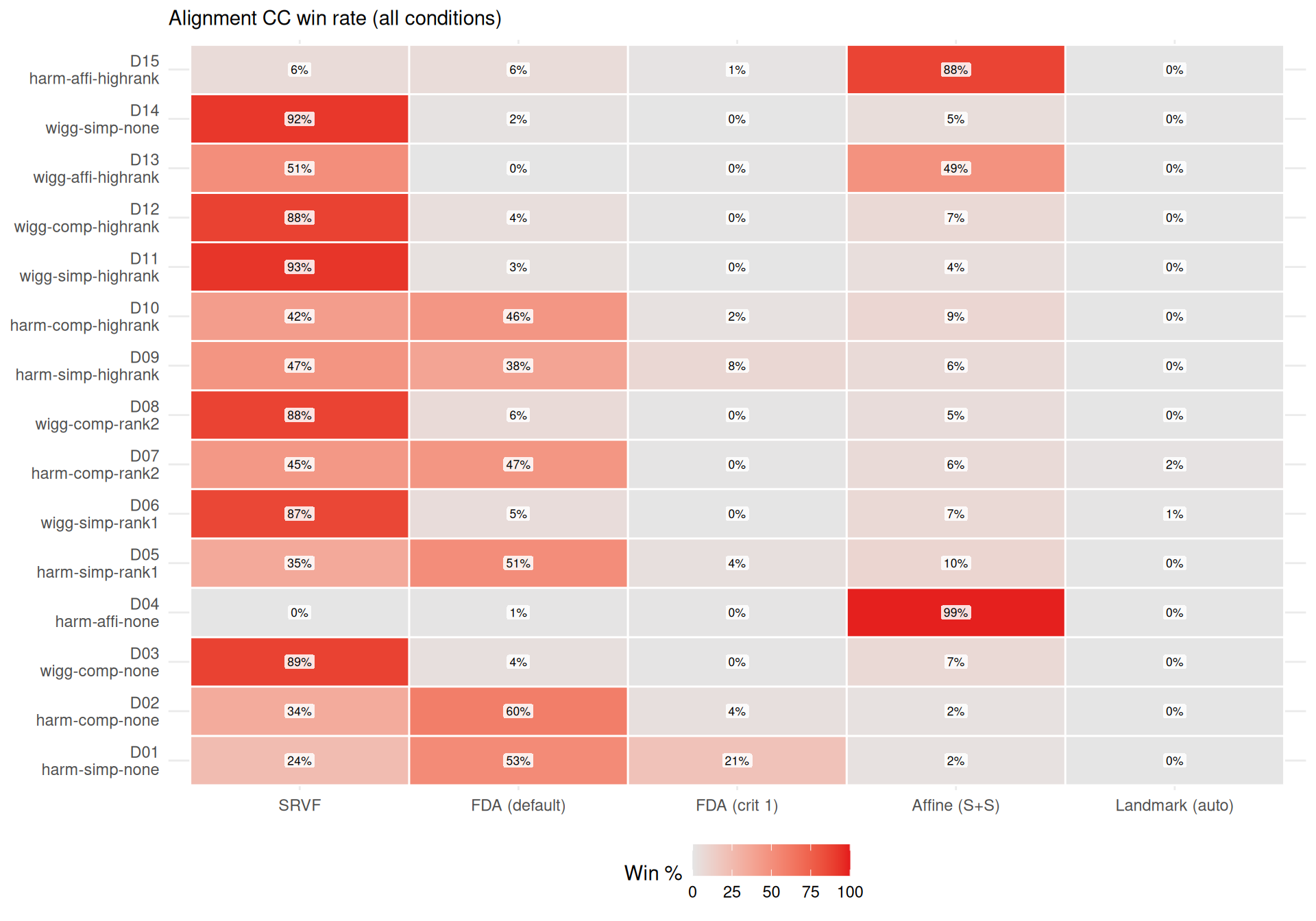
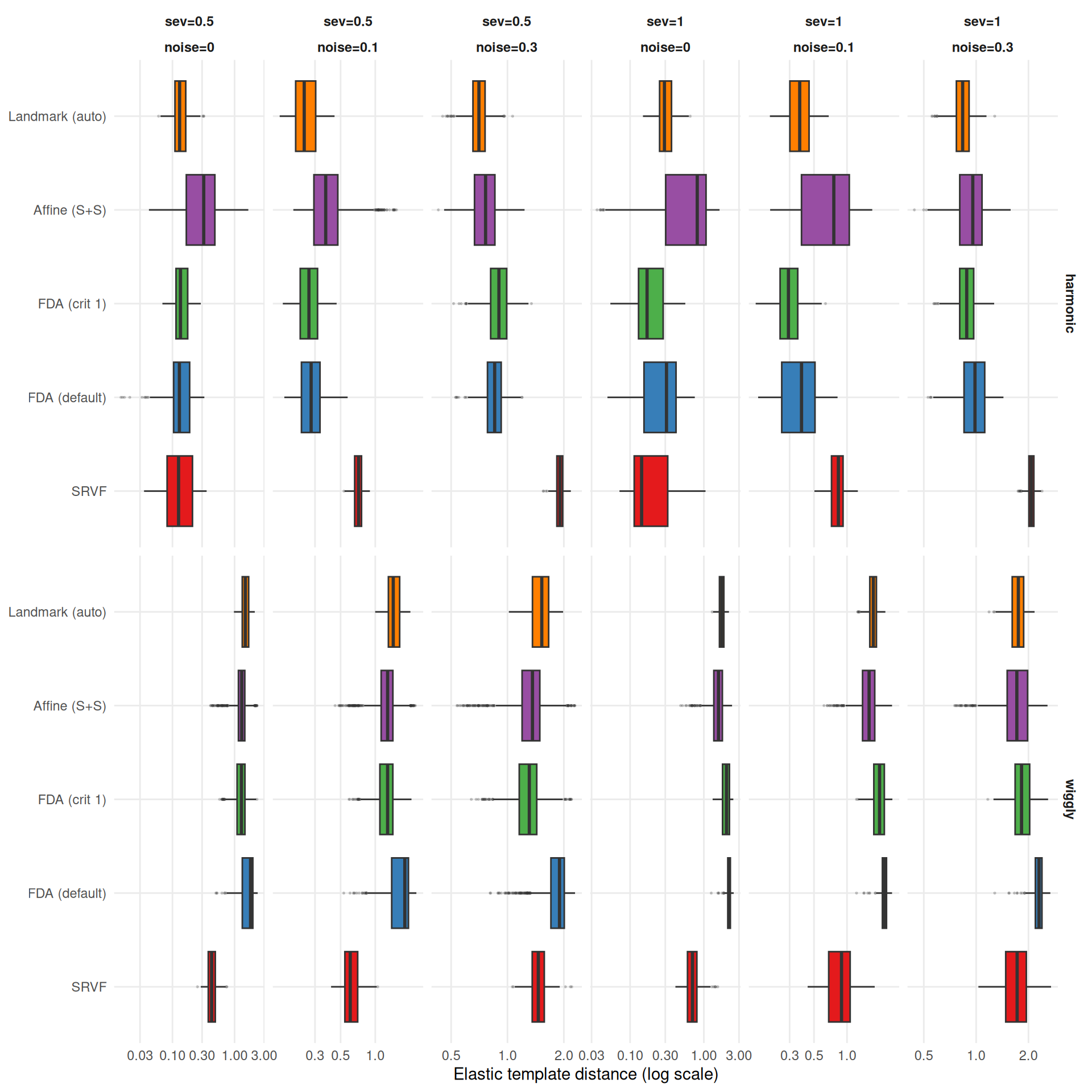
cat(sprintf(
"- **%s** wins most often on elastic distance (%s%% of conditions), suggesting it recovers template *shape* well even when its $L^2$ MISE is mediocre (phase shifts inflate MISE but not elastic distance).\n",
mlabs[wr$method[1]], wr$win_pct[1]
))
cat(sprintf(
"- **%s** rarely wins on elastic distance (%s%%), despite reasonable $L^2$ MISE — its templates have good pointwise fit but suboptimal shape recovery.\n",
mlabs[wr$method[nrow(wr)]], wr$win_pct[nrow(wr)]
))
# Noise sensitivity (template-conditioned to avoid pooling artifacts)
srvf_ratio <- wide_pooled[method == "srvf", ratio]
most_robust <- wide_pooled[which.min(ratio)]
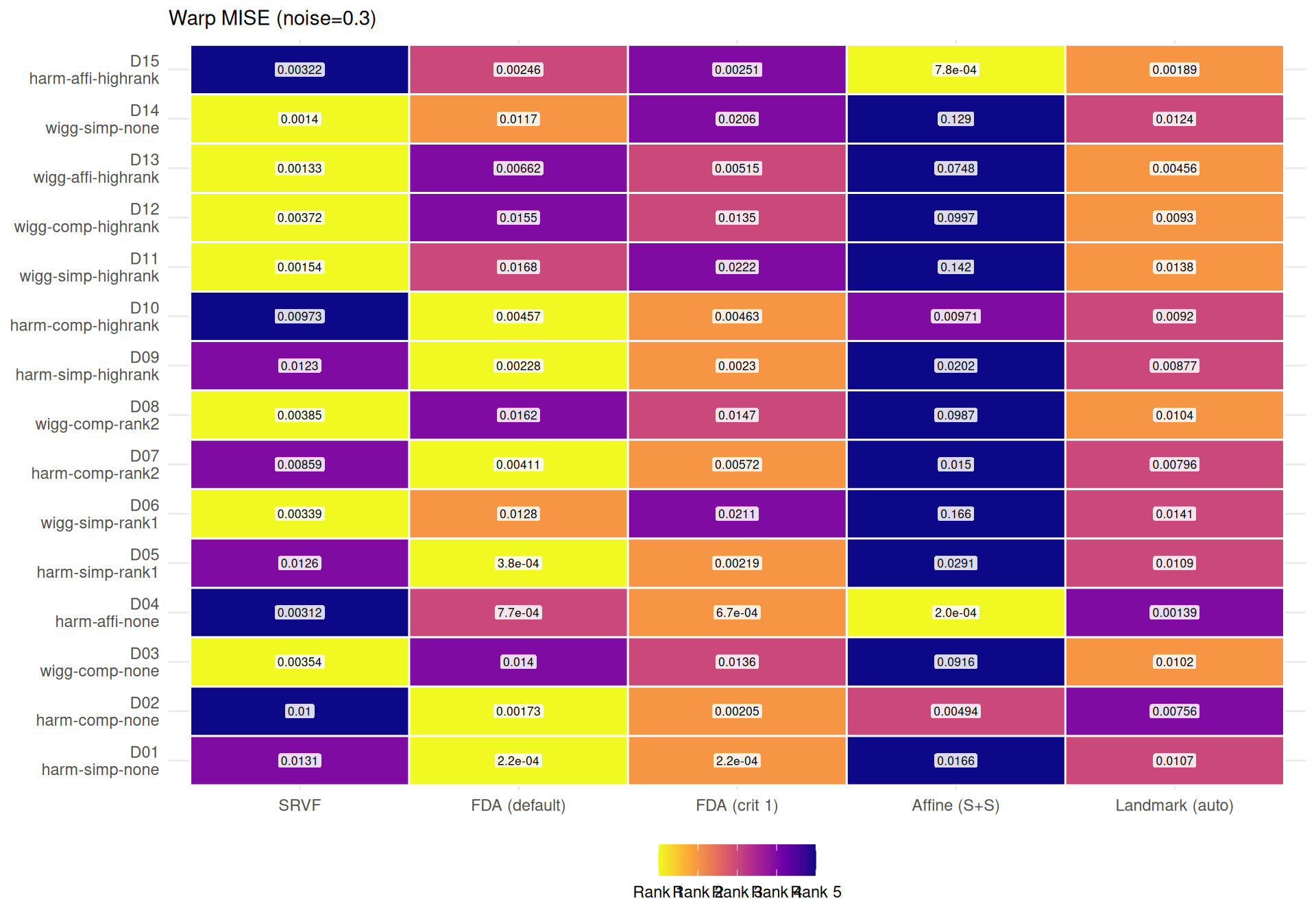
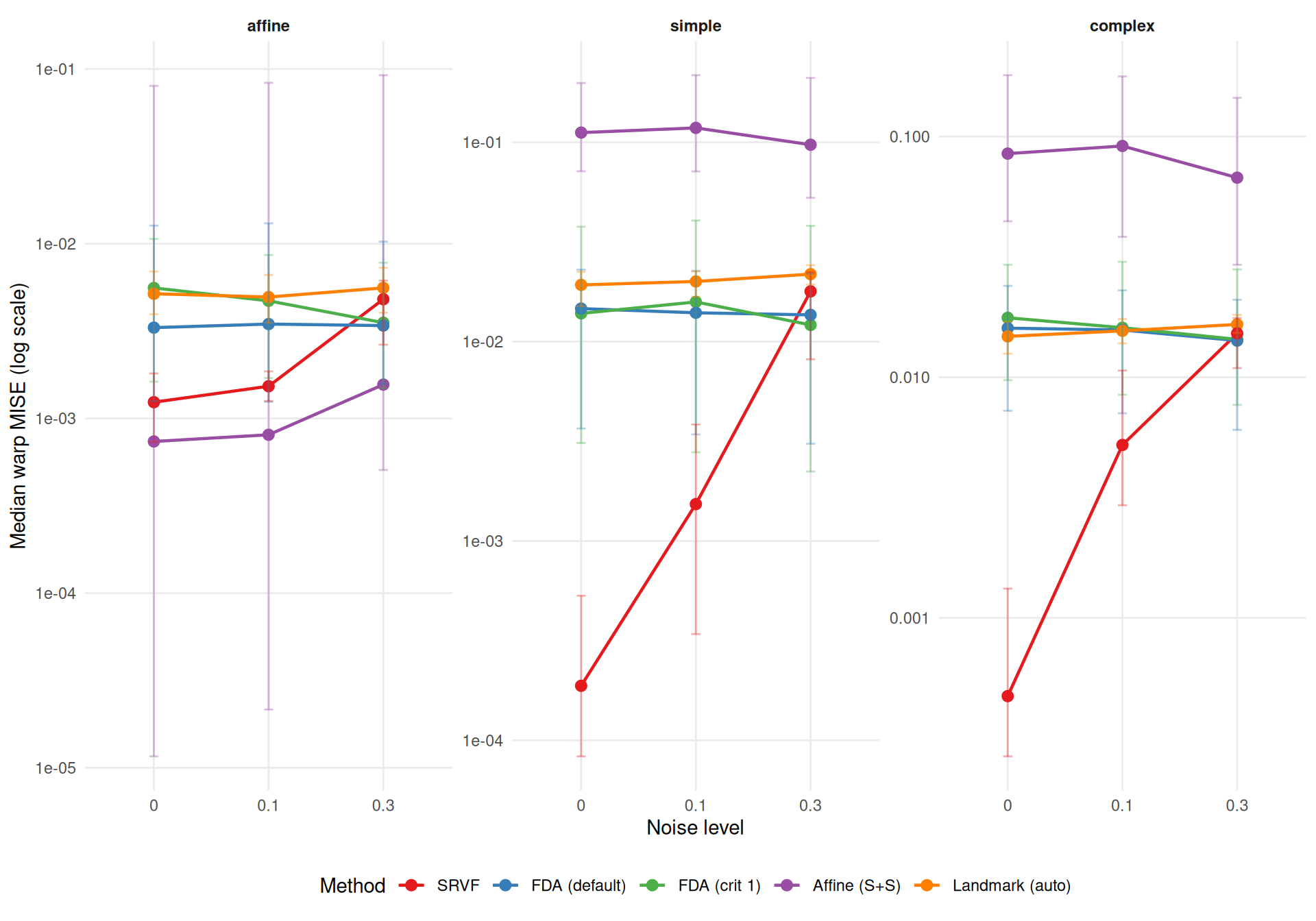
cat(sprintf(
"- **Noise sensitivity** (median ratio noise=0.3 / noise=0, conditioned on template): SRVF degrades %.1f×, while %s is most robust (%.1f×).\n",
srvf_ratio, mlabs[most_robust$method], most_robust$ratio
))
# Affine: show per-template ratios since pooling can mask heterogeneity
affine_by_tpl <- wide[method == "affine_ss"]
if (nrow(affine_by_tpl) > 0) {
affine_txt <- paste(sprintf("%s: %.1f×", affine_by_tpl$template, affine_by_tpl$ratio),
collapse = ", ")
cat(sprintf(
"- Affine (S+S) noise sensitivity varies by template (%s) — the pooled ratio masks heterogeneity. Elastic distance is always high (median %.2f–%.2f), indicating consistent template shape distortion.\n",
affine_txt,
min(noise_sens[method == "affine_ss", edist]),
max(noise_sens[method == "affine_ss", edist])
))
}
} else {
cat("*Elastic distance not available in current results.*\n")
}